BooleanNetwork¶
-
class
neet.boolean.BooleanNetwork(size, names=None, metadata=None)[source]¶ The BooleanNetwork class is a base class for all of Neet’s Boolean networks. The BooleanNetwork class inherits from both
neet.UniformNetworkandneet.boolean.SensitivityMixin, and specializes the inheritedneet.StateSpacemethods to exploit the Boolean structure.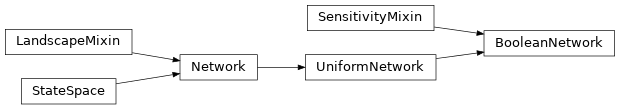
In addition to all of its inherited methods, BooleanNetwork also exposes the following methods:
subspaceGenerate all states in a given subspace. distanceCompute the Hamming distance between two states. hamming_neighborsGet all states that one unit of Hamming distance from a given state. BooleanNetwork is an abstract class, meaning it cannot be instantiated. Initialization of a BooleaNetwork requires, at a minimum, the number of nodes in the network. As with all classes that derive from
neet.Network, the user may optionally provide a list of names for the nodes of the network and a metadata dictionary for the network as a whole (e.g. citation information).Parameters: -
subspace(indices, state=None)[source]¶ Generate all states in a given subspace. This method varies each node specified by the
indiciesarray independently. The optionalstateparameter specifies the state of the non-varying states of the network. Ifstateis not provided, all nodes not inindicieswill have state0.Examples
>>> s_pombe.subspace([0]) <generator object BooleanNetwork.subspace at 0x...> >>> list(s_pombe.subspace([0])) [[0, 0, 0, 0, 0, 0, 0, 0, 0], [1, 0, 0, 0, 0, 0, 0, 0, 0]] >>> list(s_pombe.subspace([0, 3])) [[0, 0, 0, 0, 0, 0, 0, 0, 0], [1, 0, 0, 0, 0, 0, 0, 0, 0], [0, 0, 0, 1, 0, 0, 0, 0, 0], [1, 0, 0, 1, 0, 0, 0, 0, 0]]
>>> s_pombe.subspace([0], state=[0, 1, 0, 1, 0, 1, 0, 1, 0]) <generator object BooleanNetwork.subspace at 0x...> >>> list(s_pombe.subspace([0], state=[0, 1, 0, 1, 0, 1, 0, 1, 0])) [[0, 1, 0, 1, 0, 1, 0, 1, 0], [1, 1, 0, 1, 0, 1, 0, 1, 0]] >>> list(s_pombe.subspace([0, 3], state=[0, 1, 0, 1, 0, 1, 0, 1, 0])) [[0, 1, 0, 1, 0, 1, 0, 1, 0], [1, 1, 0, 1, 0, 1, 0, 1, 0], [0, 1, 0, 0, 0, 1, 0, 1, 0], [1, 1, 0, 0, 0, 1, 0, 1, 0]]
Parameters: - indicies (list, numpy.ndarray, iterable) – the indicies to vary in the subspace
- state (list, numpy.ndarray) – a state which specifes the state of the non-varying nodes
Yield: the states of the subspace
-
distance(a, b)[source]¶ Compute the Hamming distance between two states.
Examples
>>> s_pombe.distance([0, 0, 0, 0, 0, 0, 0, 0, 0], [0, 1, 0, 1, 1, 0, 1, 0, 0]) 4 >>> s_pombe.distance([0, 1, 0, 1, 1, 0, 1, 0, 0], [0, 1, 0, 1, 1, 0, 1, 0, 0]) 0
Parameters: - a (list, numpy.ndarray) – the first state
- b (list, numpy.ndarray) – the second state
Returns: the Hamming distance between the states
Raises: ValueError – if either state is not in the network’s state space
-
hamming_neighbors(state)[source]¶ Get all states that one unit of Hamming distance from a given state.
Examples
>>> s_pombe.hamming_neighbors([0, 0, 0, 0, 0, 0, 0, 0, 0]) [[1, 0, 0, 0, 0, 0, 0, 0, 0], [0, 1, 0, 0, 0, 0, 0, 0, 0], [0, 0, 1, 0, 0, 0, 0, 0, 0], [0, 0, 0, 1, 0, 0, 0, 0, 0], [0, 0, 0, 0, 1, 0, 0, 0, 0], [0, 0, 0, 0, 0, 1, 0, 0, 0], [0, 0, 0, 0, 0, 0, 1, 0, 0], [0, 0, 0, 0, 0, 0, 0, 1, 0], [0, 0, 0, 0, 0, 0, 0, 0, 1]] >>> s_pombe.hamming_neighbors([0, 1, 1, 0, 1, 0, 1, 0, 0]) [[1, 1, 1, 0, 1, 0, 1, 0, 0], [0, 0, 1, 0, 1, 0, 1, 0, 0], [0, 1, 0, 0, 1, 0, 1, 0, 0], [0, 1, 1, 1, 1, 0, 1, 0, 0], [0, 1, 1, 0, 0, 0, 1, 0, 0], [0, 1, 1, 0, 1, 1, 1, 0, 0], [0, 1, 1, 0, 1, 0, 0, 0, 0], [0, 1, 1, 0, 1, 0, 1, 1, 0], [0, 1, 1, 0, 1, 0, 1, 0, 1]]
Parameters: state (list, numpy.ndarray) – the state whose neighbors are desired Returns: a list of neighbors of the given state Raises: ValueError – if the state is not in the network’s state space
-